Most healthy people carry bacteria-infecting viruses called crassviruses in their gut, which may help to maintain the gut microbiome. However, scientists know little about this group of viruses, including their structure. This led researchers at the University of York to visit the electron Bio-Imaging Centre (eBIC) at Diamond Light Source where they captured images of these elusive viruses for the first time. Using high-voltage cryo-electron microscopes at eBIC, they overcame challenges posed by these large virus assemblies, resolving their structure to a near-atomic resolution. They found that the virus consists of a “head” region that stores viral DNA and proteins. The head is connected to a “tail” that injects these contents into bacteria, allowing virus replication to take place inside microbial cells. Additionally, they discovered a muzzle that prevents DNA and proteins from leaking out through the tail. A never-before-seen “crass” fold was spotted in the tail that the team hypothesised loosens and tightens the muzzle to control the flow of viral material into the bacterium. This study, published in Nature, offers a first glimpse at an underexplored family of widespread viruses and provides insight into their lifecycle.
The human gut is brimming with bacteria that aid with digestion and support immune health. In turn, bacteria-infecting viruses called bacteriophages keep these bacterial populations in check, preventing any one bacterial species from outgrowing the rest. Crassviruses are the most common variety of bacteriophages, accounting for nearly 95% of gut viruses in some healthy people. Oliver Bayfield, a chemist at the University of York, said, “These crassviruses and their hosts are in some sort of equilibrium that is quite stable and compatible with good gut health.” Despite their wide prevalence, when Bayfield began working on this group of viruses, scientists knew very little about them and had not determined their structure.
In early crassvirus research, scientists studied viral genomes from multiple different faecal samples in a process called “cross assembly” from which the name “crass” is derived. Initial research into this novel group of viruses has largely been restricted to studying their genes as a result. Now, Bayfield and his colleagues have for the first time observed what these microbes look like under the microscope in granular detail. In collaboration with eBIC, they used cryo-electron microscopy (cryoEM) to solve the structure of ΦcrAss001, the first crassvirus ever isolated from human faecal samples.
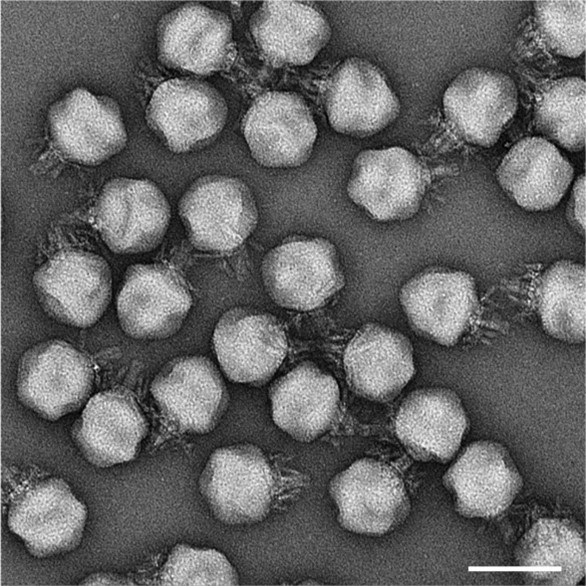
CryoEM proves valuable for studying protein complexes, but extremely large protein assemblies like crassviruses come with their own challenges. To resolve the structure of any protein complex, one needs to capture hundreds or thousands of examples and average them together. However, because the crassvirus is so large, each image possessed fewer particles than usual for a cryoEM experiment. The eBIC team helped Bayfield capture enough copies of the virus by zooming out to a lower magnification.
Bayfield added that he was able to produce a high resolution cryoEM structure in part thanks to eBIC’s high-voltage instruments. he explained,
For large samples you need to have 300kV in order to compensate for the thickness of the sample.
Bayfield and his colleagues reconstructed the virus’ anatomy to a near-atomic resolution of 3.1 Ångströms (Å). Despite crassviruses sharing very few of their DNA sequences with other known bacteriophages, Bayfield noted that they adopted similar, conserved structures. FcrAss001 possesses an icosahedral head and a tail, like other viruses, such as the famous moon-lander shaped T4 bacteriophage that infects E. coli. These anatomical parts serve crucial functions for the virus: the head stows away viral DNA in neatly folded piles while the tail threads the DNA out of the head and into a bacterial host.
The head mostly contained neatly and tightly packed DNA without much free space. However, the team spotted a hollow region near the base of the head where DNA was sparse and in disarray. They hypothesised that viral proteins might occupy the seemingly vacant space. Bayfield said,
We had some indication that these proteins were carried along with the virion from proteomics data.
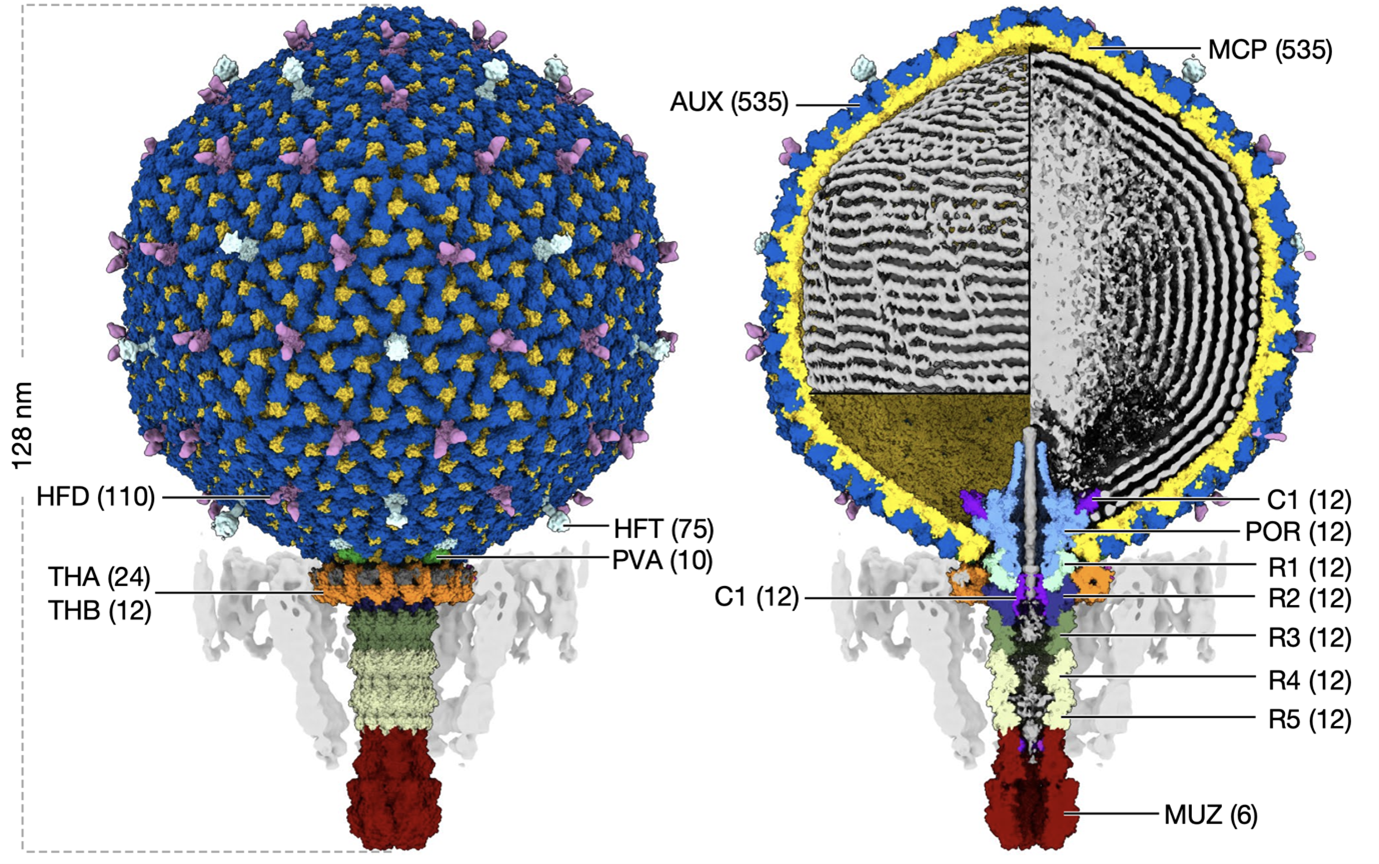
The team estimated how spacious these DNA-free regions were, calculating that the virus could fit up to six copies of the relatively large RNA polymerase complexes or 12 copies of smaller viral proteins. Bayfield explained that the proteins “seem to be stored in a fairly flexible or disordered manner, and that’s likely because that would help them get out of the virus and into the cell.”
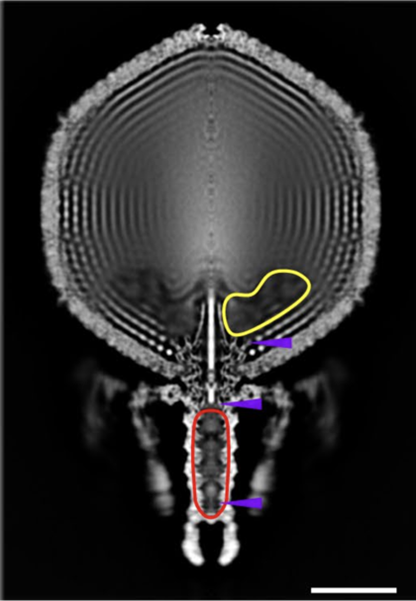
Once a crassvirus finds a bacterial host, it docks at its surface using its tail and injects its contents into the microbe. The researchers estimated that the capsid stores DNA and proteins in the head at 100 atmospheres, suggesting the high pressure might trigger the contents to burst out through the tail, like an overfilled, untied water balloon. How then does the virus prevent its DNA and protein from leaking out before it encounters a bacterium?
Bayfield and his colleagues found that tip of tail is capped with an assemblage of six proteins that form a muzzle. They also discovered a novel protein fold in the virus’ tail that they name the “crass fold”, which they hypothesise could tighten or loosen the muzzle through conformational changes, thereby controlling the outpour of viral DNA and proteins.
ΦcrAss001 is just one of a multitude of crassviruses. In future work, Bayfield aims to solve the structure of different viral species, including close and distant relatives, so he can triangulate what it means to be “crass” in the viral world.
To find out more about eBIC or discuss potential applications, please contact eBIC director Peijun Zhang: [email protected].
This work was supported by the Wellcome Trust (grant 206377, 224665 to A.A.A.), Instruct-ERIC (PID: 10175), and Diamond Light Source for access to eBIC under proposal EM19832 funded by the Wellcome Trust, MRC and BBRSC. The cryo-EM facility at the University of York was supported by the Wellcome Trust, grant 206161. A.N.S. was supported by the Wellcome Trust (grant 220646) and ERC (grant agreement no. 101001684). E. V. Khokhlova and C.H. were supported by Science Foundation Ireland (SFI) under Grant no. SFI/12/RC/2273. N.Y. and E. V. Koonin are supported by the Intramural Research Program of the National Institutes of Health of the USA (National Library of Medicine). Molecular graphics and analyses were performed with UCSF ChimeraX, developed by the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco, with support from National Institutes of Health R01-GM129325 and the Office of Cyber Infrastructure and Computational Biology, National Institute of Allergy and Infectious Diseases.
Bayfield OW, Shkoporov AN, Yutin N, Khokhlova EV, Smith JLR, Hawkins DEDP, Koonin EV, Hill C and Antson AA. Structural atlas of a human gut crassvirus. Nature 617 (7959), 409–416 (2023). doi: 10.1038/s41586-023-06019-2
Diamond Light Source is the UK's national synchrotron science facility, located at the Harwell Science and Innovation Campus in Oxfordshire.
Copyright © 2022 Diamond Light Source
Diamond Light Source Ltd
Diamond House
Harwell Science & Innovation Campus
Didcot
Oxfordshire
OX11 0DE
Diamond Light Source® and the Diamond logo are registered trademarks of Diamond Light Source Ltd
Registered in England and Wales at Diamond House, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, United Kingdom. Company number: 4375679. VAT number: 287 461 957. Economic Operators Registration and Identification (EORI) number: GB287461957003.